SPANDx: a genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets, BMC Research Notes
By A Mystery Man Writer
Description

Frontiers DIVIS: Integrated and Customizable Pipeline for Cancer Genome Sequencing Analysis and Interpretation

Dynamics of genetic variation in Transcription Factors and its implications for the evolution of regulatory networks in Bacteria
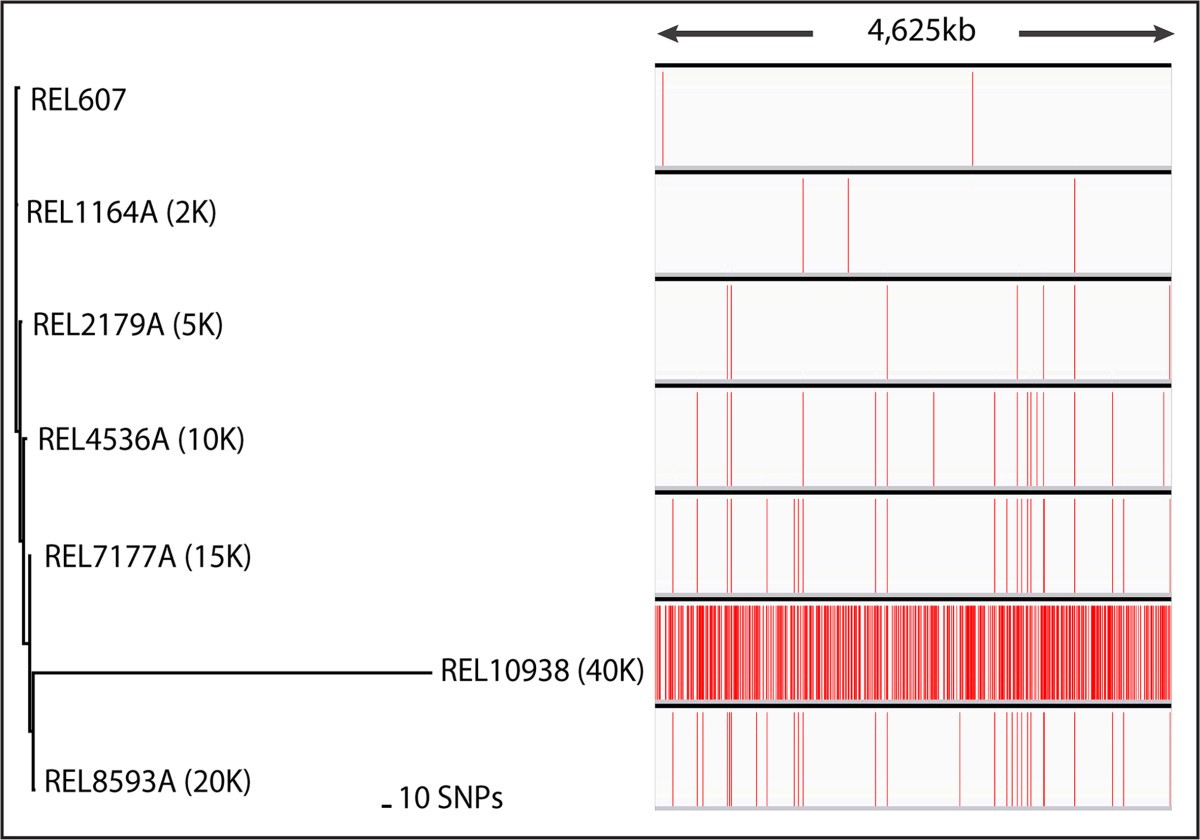
SPANDx: a genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets, BMC Research Notes
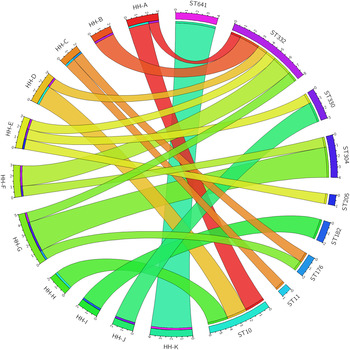
Whole genome sequencing reveals extensive community-level transmission of group A Streptococcus in remote communities, Epidemiology & Infection

Keeping up with the pathogens: Improved antimicrobial resistance detection and prediction from Pseudomonas aeruginosa genomes

Whole-Genome Sequencing of Burkholderia pseudomallei Isolates from an Unusual Melioidosis Case Identifies a Polyclonal Infection with the Same Multilocus Sequence Type
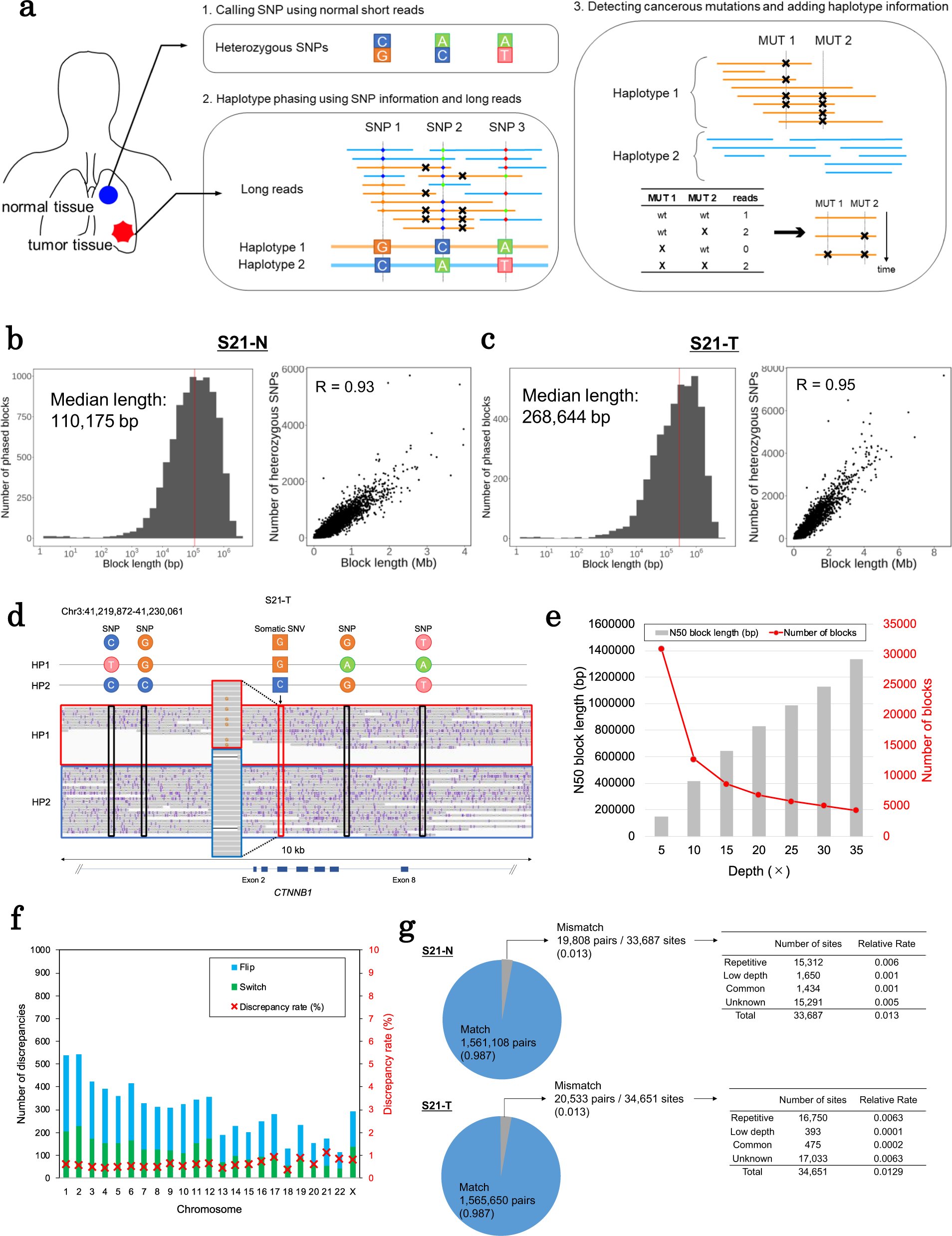
Phasing analysis of lung cancer genomes using a long read sequencer

Comparison of two African rice species through a new pan-genomic approach on massive data
Opportunistic pathogens and large microbial diversity detected in source-to-distribution drinking water of three remote communities in Northern Australia

Metabolism of ʟ -arabinose converges with virulence regulation to promote enteric pathogen fitness
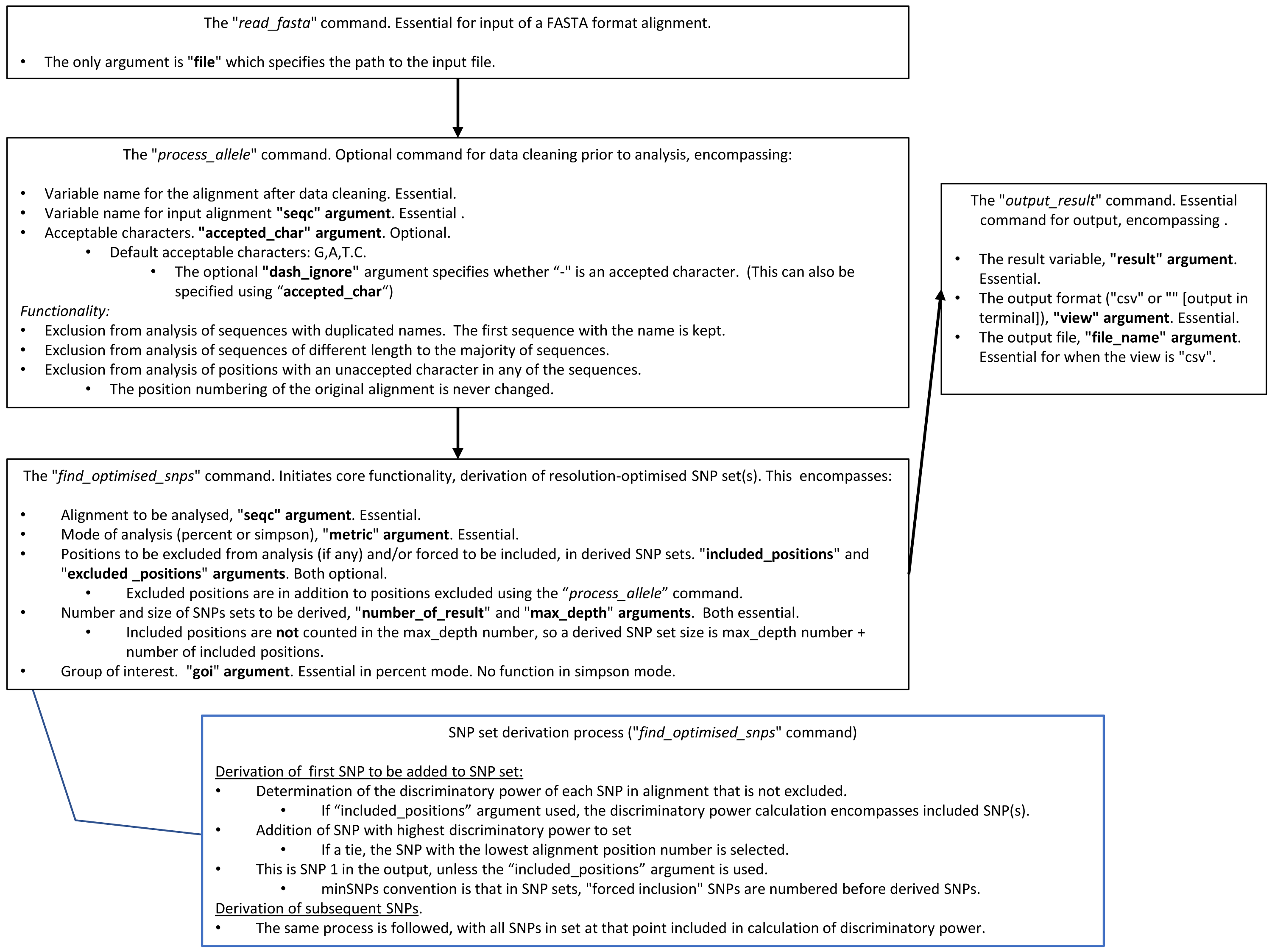
minSNPs: an R package for the derivation of resolution-optimised SNP sets from microbial genomic data [PeerJ]

The Northern Arizona SNP Pipeline (NASP): accurate, flexible, and rapid identification of SNPs in WGS datasets

Can non-typeable Haemophilus influenzae carriage surveillance data infer antimicrobial resistance associated with otitis media?
from
per adult (price varies by group size)







