Technical spotlight: Detecting small- and medium-length copy number variants by whole-genome sequencing
By A Mystery Man Writer
Description
Historically, detecting different sizes of genetic variants has required using multiple different tests. By combining Illumina WGS with secondary analysis algorithms built into the DRAGEN Bio-IT Platform, researchers can achieve high-sensitivity detection of all these different variant types using a mixture of methods described here.

The era of whole genome sequencing : Revista Pesquisa Fapesp
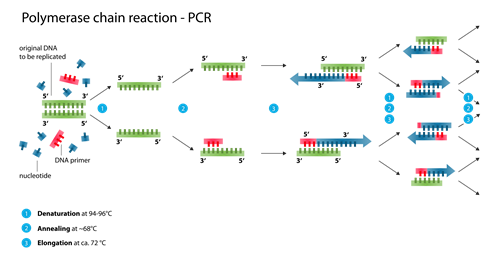
Amplicon Sequencing

Rami Mehio on LinkedIn: Edico Genome's team that built DRAGEN in

Steven Barnard posted on LinkedIn

An Overview of Next-Generation Sequencing

Samuel Strom, PhD FACMG on LinkedIn: whoa. I mean I knew thing

Genomics Articles Recent genomics discoveries by Illumina scientists

DRAGEN 4.2 release: CNV calling on genes

Copy number variation detection in whole-genome sequencing data using the Bayesian information criterion
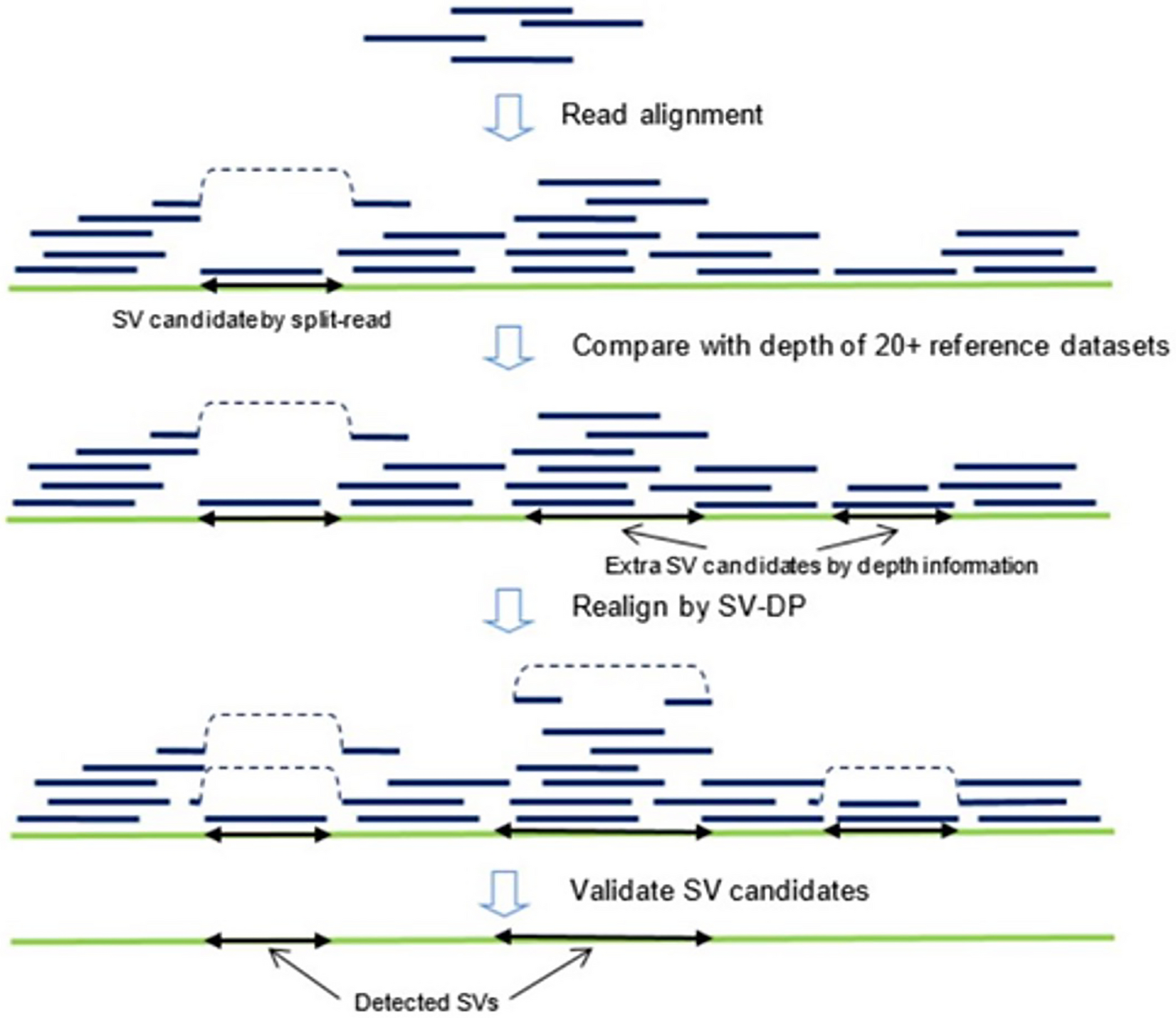
Detecting structural variations with precise breakpoints using low-depth WGS data from a single oxford nanopore MinION flowcell

Identification of Copy Number Alterations from Next-Generation Sequencing Data

Detecting copy number variation in next generation sequencing data from diagnostic gene panels, BMC Medical Genomics

Rami Mehio على LinkedIn: Using whole-genome sequencing to evaluate
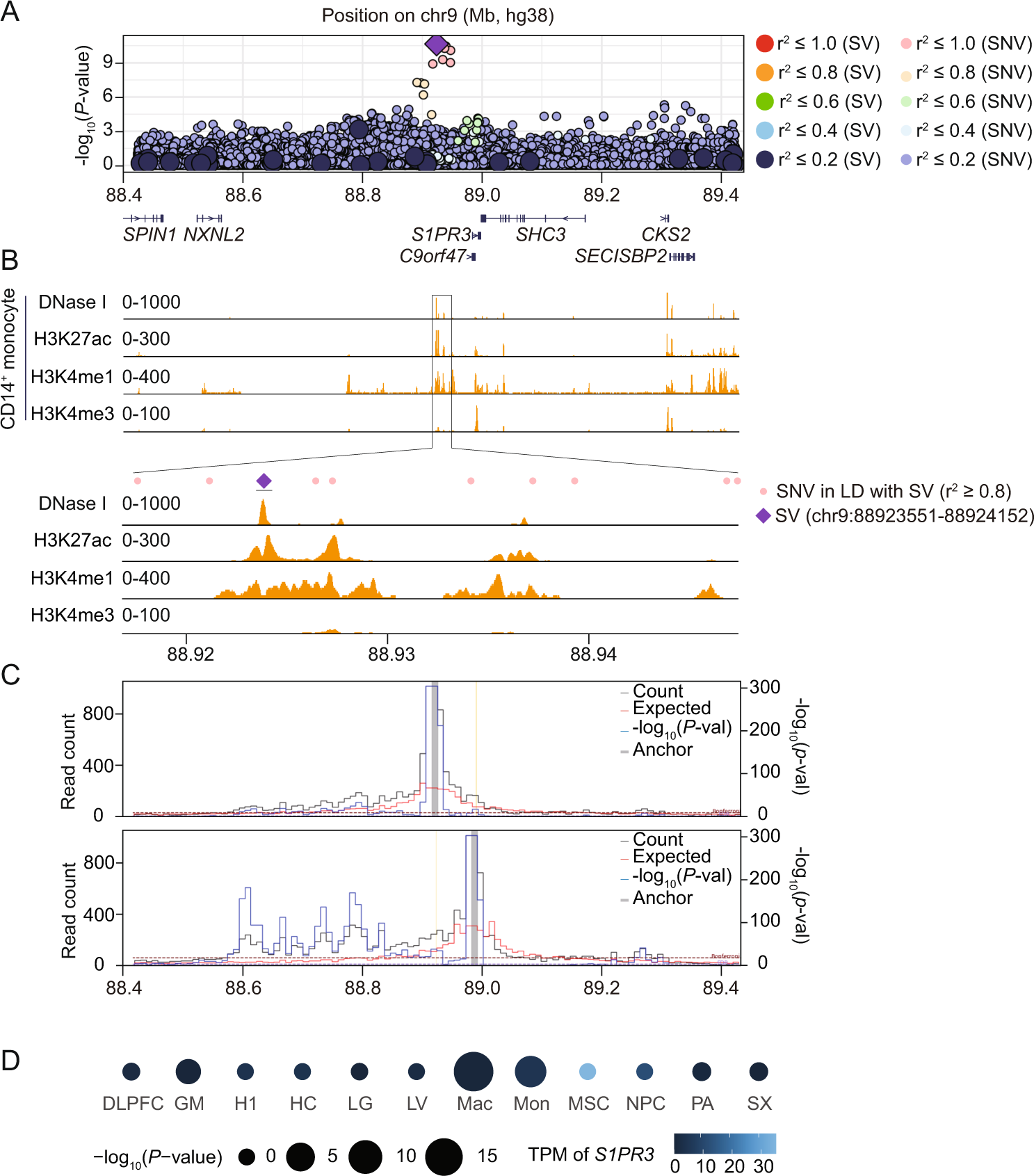
Whole genome sequencing identifies structural variants contributing to hematologic traits in the NHLBI TOPMed program
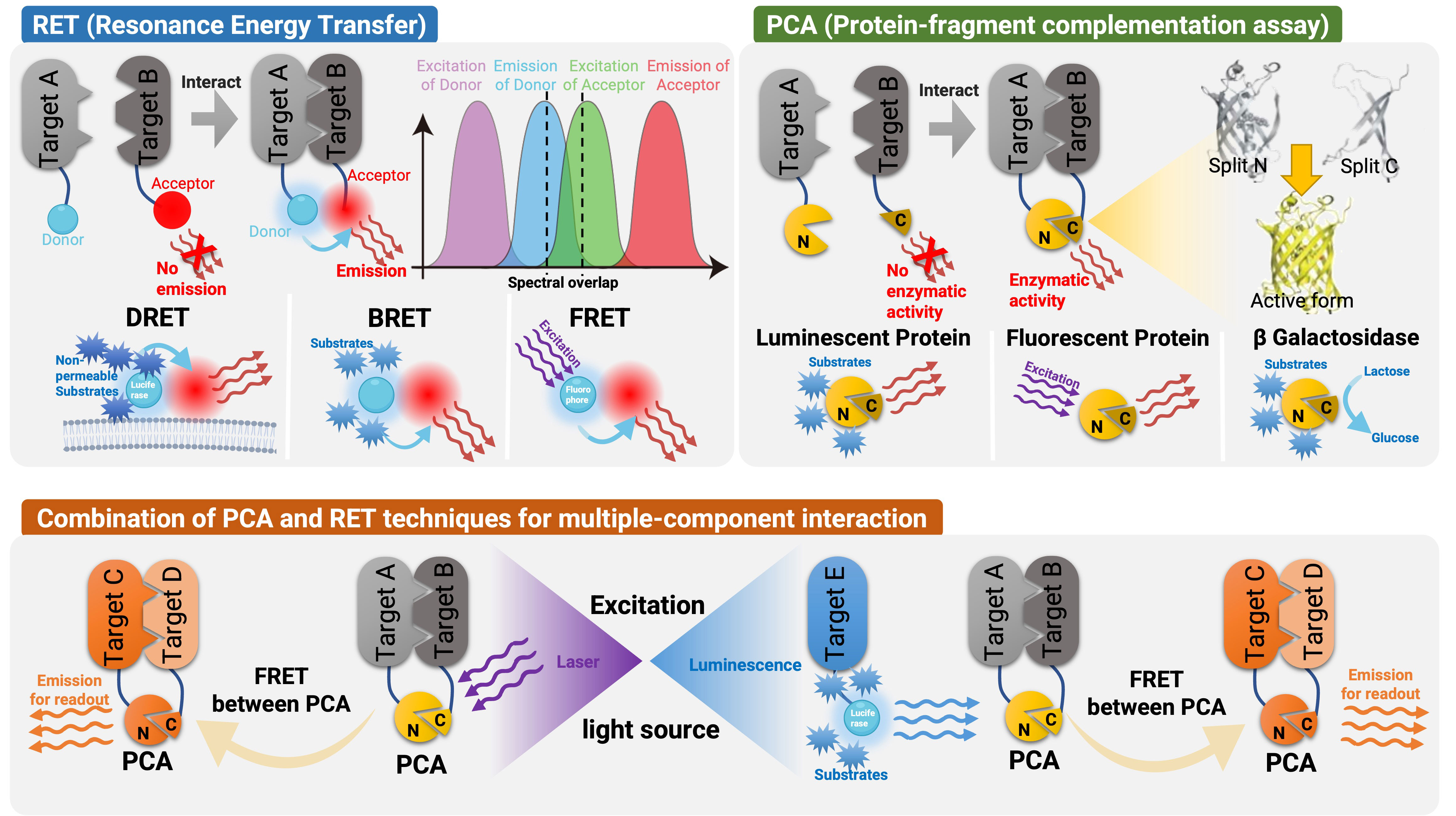
Frontiers Detecting and measuring of GPCR signaling – comparison of human induced pluripotent stem cells and immortal cell lines
from
per adult (price varies by group size)






